Integrating Machine Learning and Genomic Data to Study Microbiome-Associated SNPs in Beef Cattle
DOI:
https://doi.org/10.65718/inspireHealth.2026.2010Keywords:
Microbiome-SNP Interactions, Beef Cattle Genomics, Computational Biology, LASSO Algorithm, Genomic Selection, Host-Microbiome DynamicsAbstract
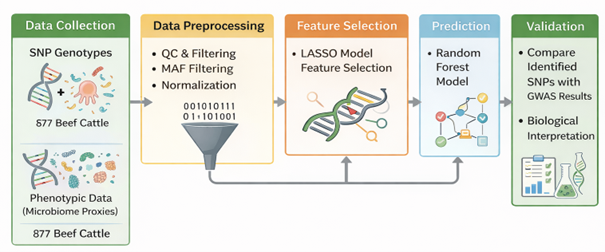
The relationship between the microbiome and single-nucleotide polymorphisms (SNPs) in beef cattle could help improve animal health, productivity, and sustainability. In this study, we use computational analysis to exam-ine how the microbiome and SNPs are connected. The primary objective is to identify significant associations that could inform breeding strategies and disease management practices within the context of beef cattle pro-duction. We use computational biology techniques, focusing on feature selection methods such as the LASSO (Least Absolute Shrinkage and Selection Operator) algorithm, to identify important SNPs associated with micro-bial composition. By combining genomic and microbiome data, we aim to clarify the interactions that affect the health and productivity of cattle. This research adds to our understanding of how hosts and their microbiomes interact, which could help us develop more personalized methods for managing livestock in the future.
Downloads
Published
Issue
Section
Categories
License
Copyright (c) 2026 Inspire Health Journal

This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License.



